- Blog
- Pling rabat
- Cronometer app reviews
- Jamie raskin wikipedia
- Running python in rstudio
- Nuclear throne platforms
- Scribus for mac edit text
- Ernest hemingway editor
- Colormunki display vs spyder 5 pro
- Xml tools for excel
- Wirecast pro multiple outputs
- Altered consciousness
- Uk driving test pass rate first time
- Speech timer estimator
- Mac media player speed control
- Pacesetter directional drilling
- Blog
- Pling rabat
- Cronometer app reviews
- Jamie raskin wikipedia
- Running python in rstudio
- Nuclear throne platforms
- Scribus for mac edit text
- Ernest hemingway editor
- Colormunki display vs spyder 5 pro
- Xml tools for excel
- Wirecast pro multiple outputs
- Altered consciousness
- Uk driving test pass rate first time
- Speech timer estimator
- Mac media player speed control
- Pacesetter directional drilling
- #RUNNING PYTHON IN RSTUDIO HOW TO#
- #RUNNING PYTHON IN RSTUDIO CODE#
- #RUNNING PYTHON IN RSTUDIO DOWNLOAD#
#RUNNING PYTHON IN RSTUDIO CODE#
You can execute code from Python scripts line-by-line using the Run button (or Ctrl+Enter) in the same way as you execute R code line-by-line.
#RUNNING PYTHON IN RSTUDIO HOW TO#
See the article on the reticulate R Markdown Python Engine for full details on using Python chunks within R Markdown documents, including how to call Python code from R chunks and vice-versa. R Notebooks can also display matplotlib plots inline when they are printed from Python chunks: The article on Calling Python from R describes the various ways to access Python objects from R as well as functions available for more advanced interactions and conversion behavior. from Pandas data frame to R data frame or NumPy 2D array to R matrix). R and Python objects are also shared across languages with conversions done automatically when required (e.g. Python objects all exist in a single persistent session so are usable across chunks just like R objects. For example, here we use pandas to do some data manipulation then plot the results with ggplot2: R Notebooks have been enhanced to support executing Python chunks using the reticulate Python engine. Īll of the features described below require that you have previously installed the reticulate package, which you can do as follows:
#RUNNING PYTHON IN RSTUDIO DOWNLOAD#
You can download the RStudio v1.2 preview release here. However, if you are using reticulated Python within an R project then RStudio provides a set of tools that we think you will find very useful. Note that for data science projects that are Python-only, we still recommend IDEs optimized for that, such as JupyterLab, P圜harm, Visual Studio Code, Rodeo, and Spyder. Sourcing Python scripts using the reticulate source_python() function.Ĭode completion and inline help for Python. Line-by-line execution of Python code using the reticulate repl_python() function. Support for executing reticulated Python chunks within R Notebooks.ĭisplay of matplotlib plots within both notebook and console execution modes. New features in RStudio v1.2 related to reticulate include: If you are an R developer that uses Python for some of your work or a member of data science team that uses both languages, reticulate can dramatically streamline your workflow. The reticulate package makes it possible to embed a Python session within an R process, allowing you to import Python modules and call their functions directly from R. Today we’re taking a look at enhancements we’ve made around the reticulate package (an R interface to Python). Last week on the blog we talked about new features for working with SQL and D3. The import () function can be used to import any Python module.One of the primary focuses of RStudio v1.2 is improved support for other languages frequently used with R. You can call methods and access properties of the object just as if it was an instance of an R reference class. Types are converted as follows: If a Python object of a custom class is returned then an R reference to that object is returned. How does import function in Python convert to your reference? f = open(“test.txt”, ‘w’) Every time when we open the file, as a good practice we need to ensure to close the file, In python, we can use close () function to close the file. In order to write the data into a file, we need to open the file in write mode. In order to read the file, first, we need to open the file in reading mode. Python provides a built-in function called open () to open a file, and this function returns a file object called the handle and it is used to read or modify the file.


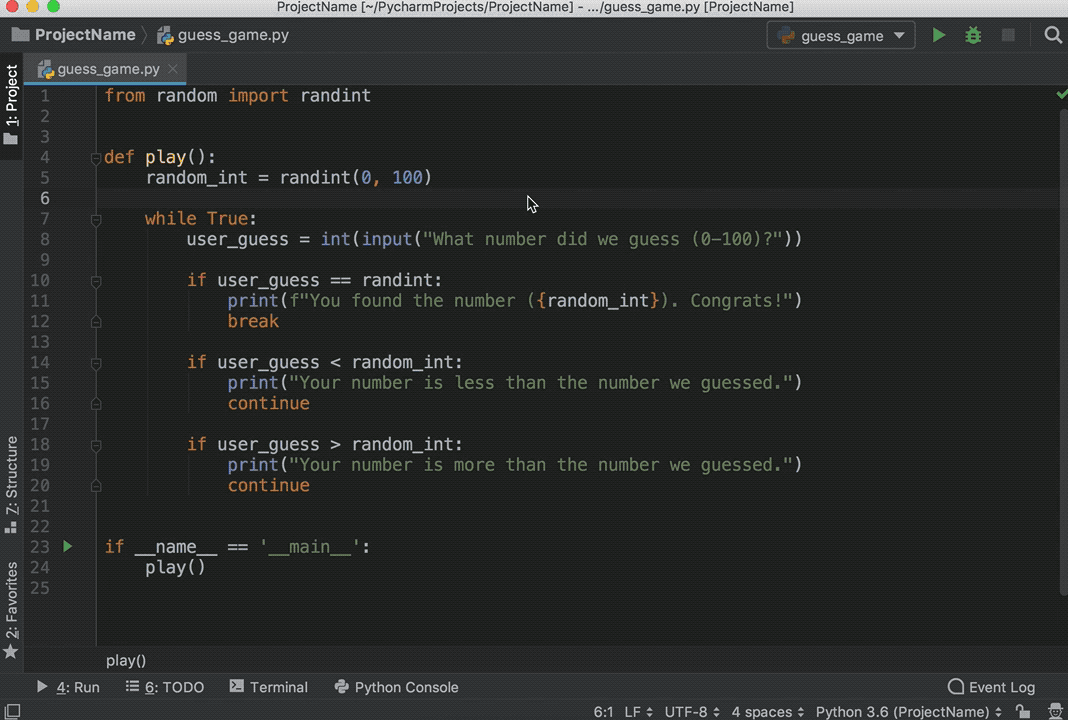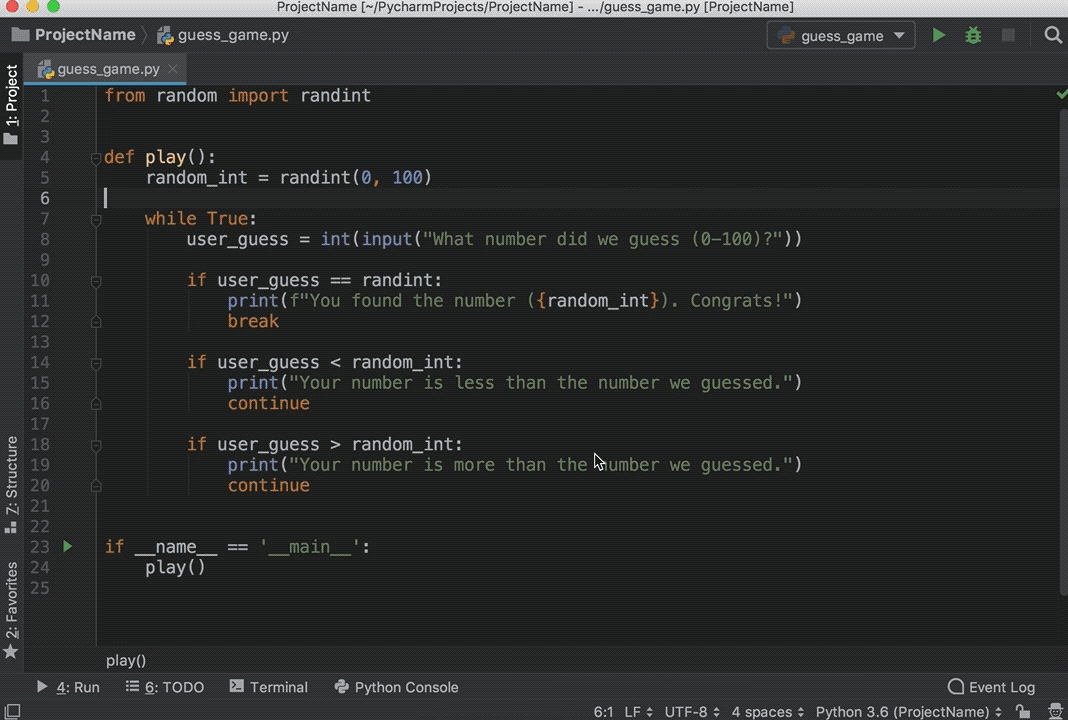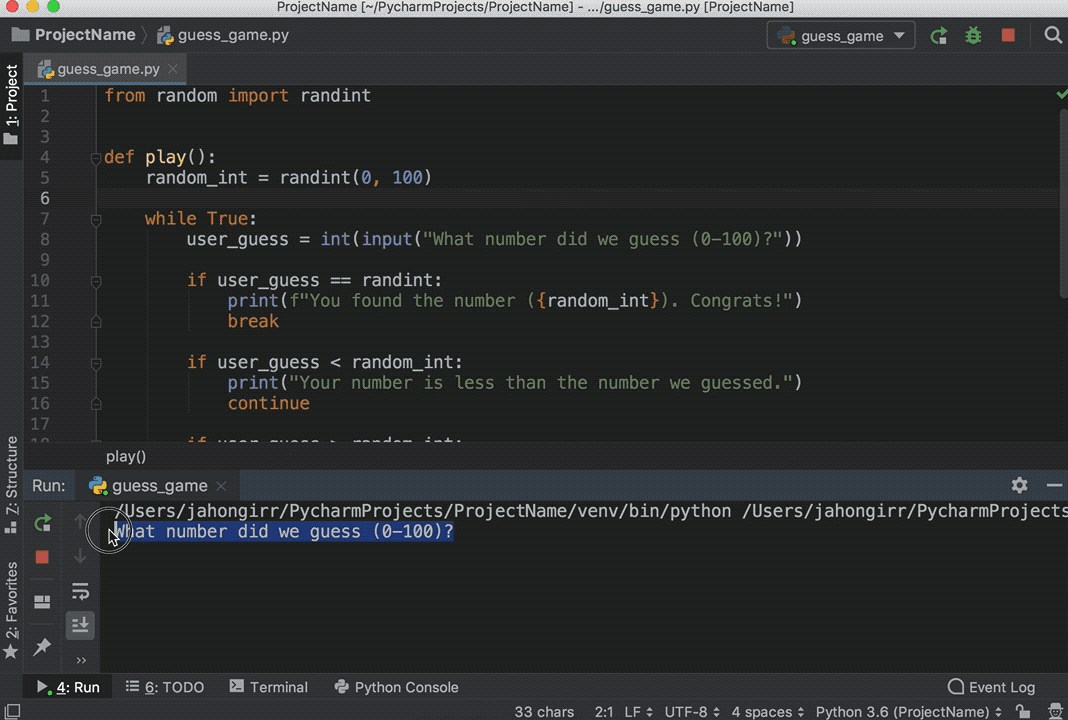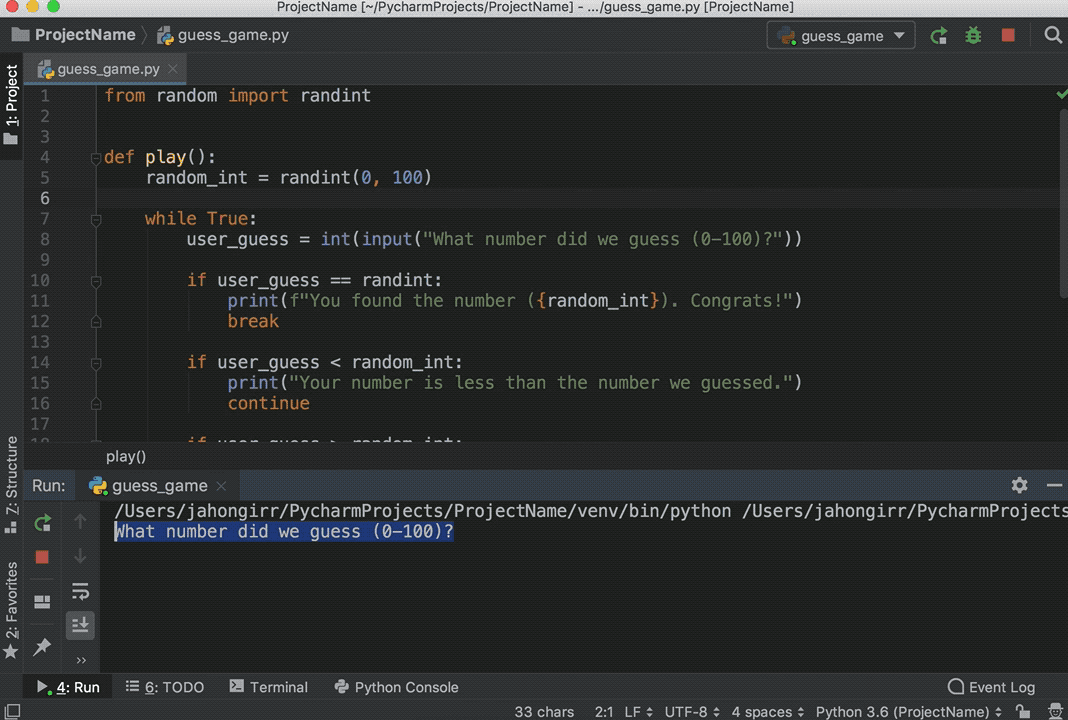
A file is a named location on the disk which is used to store the data permanently. It embeds a Python session within an R session, and allows you to pass objects between the two sessions.īut in Python 3 the raw_input () function was removed and renamed to input (). With reticulate you can run your Python scripts in RStudio. Any objects created within the Python session are available in the R session via the py object.
